Publishing over 100 genomes: an important milestone for the Tree of Life programme
| 19 May, 2022 | Abbie nicholson |

We have published our first 100 genomes!
Over 100 high-quality reference genome sequences are now available in our Tree of Life Gateway. We would like to thank everyone involved in this amazing effort!
Read on for more details about the project and three highlights from the Gateway so far.
The Tree of Life programme
What it’s all about
In May 2021, Wellcome Open Research launched the Tree of Life Gateway. This Gateway is an open publishing hub for the Darwin Tree of Life, Aquatic Symbiosis Genomics, and 25 Genomes for 25 years projects, all supported by the Wellcome Sanger Institute.
The main goal of the project is to publish the genome sequences of thousands of species living in Britain and Ireland as fast as possible. Rapid dissemination can help make these sequences discoverable and reusable for researchers worldwide, enabling progress and further research in the field of genomics.
A few words from Mark Blaxter
“At the heart of this project to sequence all life is an ambition to make our genomic data available as openly and rapidly as possible. We release assemblies and their annotations as soon as we generate them, announcing them via the Genome Notes published on our gateway at Wellcome Open Research. These short, definitive publications contain the key specimen metadata and assembly metrics. They also ensure that credit accrues directly to those involved in sequencing these genomes: from collectors and taxonomists in the field, through those responsible for DNA barcoding to the teams extracting, sequencing and assembling chromosomal-level genome assemblies.
“This method of rapid, open publication allows researchers across the globe to exploit our data immediately, assured that it is free to use. We expect our assemblies to have significant impact for scientists focused on evolution, ecology and conservation, as well as those developing new biotechnologies and biomedicines. Even at this early and thrilling milestone of 100-plus Genome Notes, we see our data fuelling all kinds of investigation. In short, this rapid and open output is already transforming the way we do biology.”
Professor Mark Blaxter, Programme Lead for the Tree of Life Programme at the Wellcome Sanger Institute
Fast processes
The Platform’s key strengths include speed of article acceptance, publication, and passing peer review – but for genome sequences in the Tree of Life Gateway, these processes take place even faster. The genomes are sequenced rapidly through machine learning, and then become ready for immediate publication.
The Platform’s post-publication peer review model further accelerates the timeline for potential impact, as genomics researchers can access and use the genome sequences in their own projects without the delays of more traditional peer review.
Table 1 shows that most genome sequences are accepted 13 days after submission. The equivalent for Wellcome Open Research overall is 29 days. The median time from acceptance to publication is 10 days for the Platform and 9 days for the Gateway. Sequences on average pass peer review 36 days after publication. For Wellcome Open Research articles in general this takes around 75 days.

Three highlights from the Gateway
With over 100 genomes published in the Tree of Life Gateway since its launch in 2021, browsing the Gateway offers a fascinating insight into the wide range of species found across Britain and Ireland. Let’s take a look at three highlights…
- Eimeria tenella is a common gut parasite that can affect chickens’ welfare, with great implications for the poultry industry. It is the first single-celled protist to have its genome published within the context of the Tree of Life programme.
- The large tortoiseshell (Nymphalis polychloros) is a type of butterfly from the insect order Lepidoptera. This species has made a mysterious reappearance in the United Kingdom after becoming extinct in the 1950s. The genome may help explore the butterfly’s origins, behaviors as an adult or caterpillar, and reasons for its disappearance.
- The common toad (Bufo Bufo) is an amphibian whose populations in Britain have declined in recent decades. Sequencing this genome can help identify physical and evolutionary differences between the common toad and the common frog – another amphibian sequenced within the Tree of Life programme – as well as compare populations of toads that are thriving to those that are dying out, to help aid conservation.
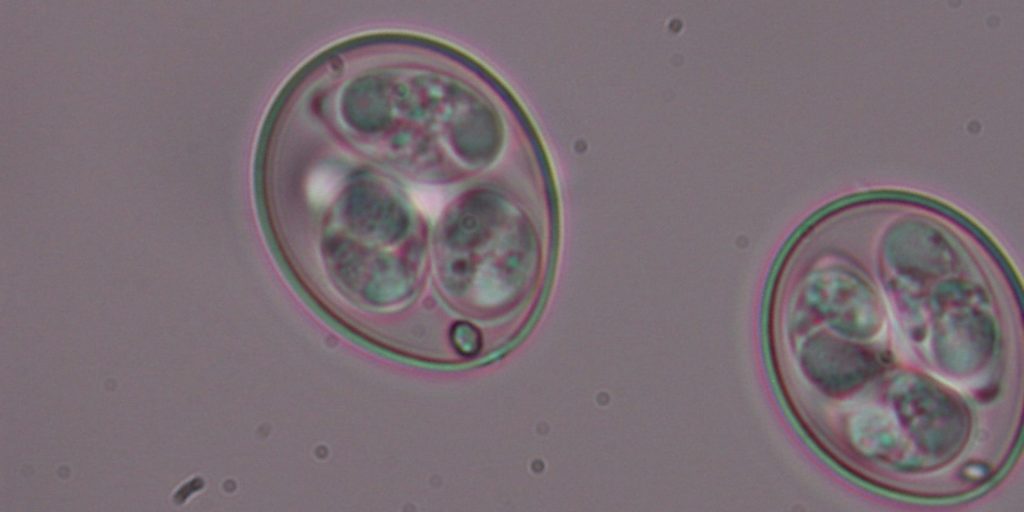
Eimeria tenella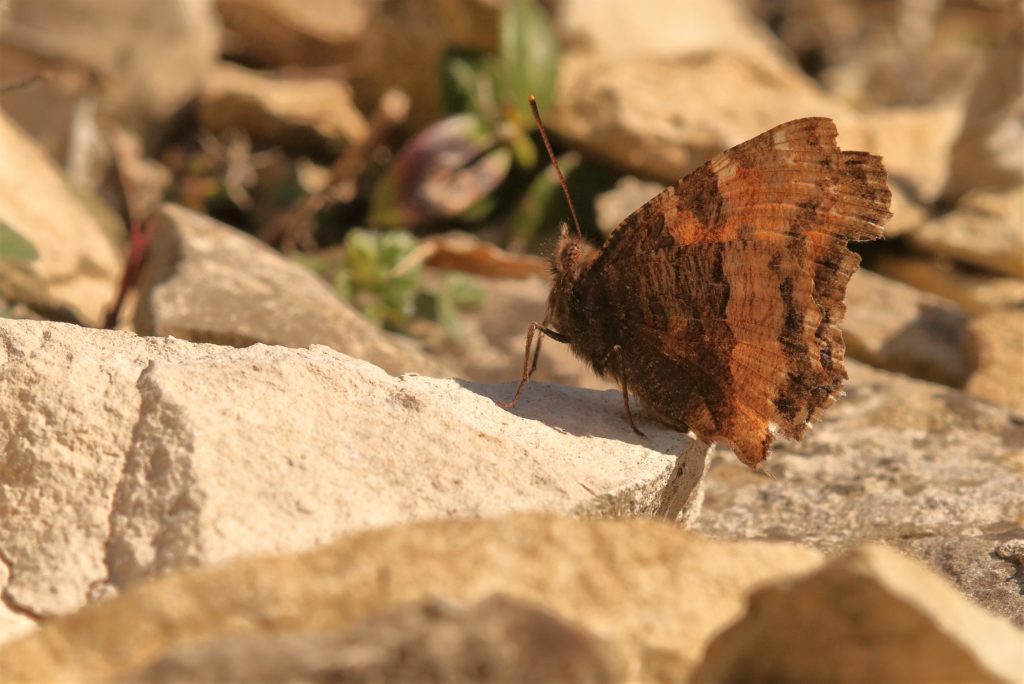 
Nymphalis polychloros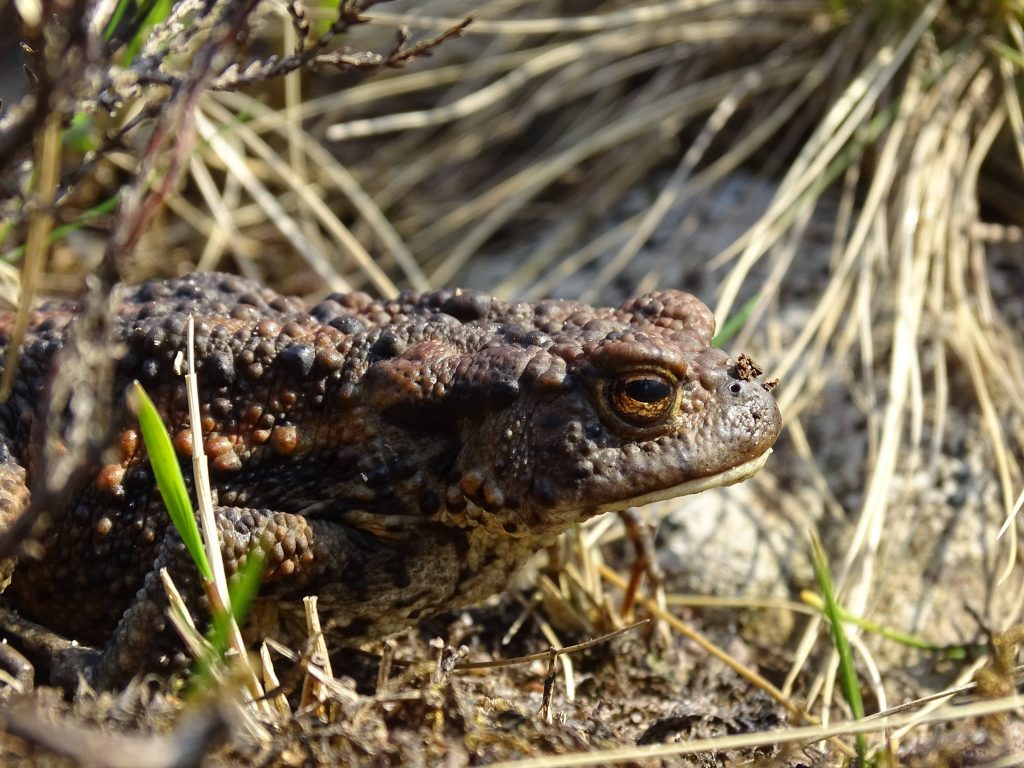 
Bufo bufo
How can you support the Tree of Life?
We invite all genomics researchers to contribute to the Tree of Life programme as community reviewers, by assessing the genome sequences published in the Tree of Life Gateway.
This is a unique opportunity for reviewers to join an innovative project and receive credit for their work thanks to Wellcome Open Research’s open peer review process. This initiative is ideal for early career researchers and PhD students who are looking to get started with peer review and build connections with the wider research community.
If you are interested in joining the Tree of Life community reviewer group, please sign up here, or contact us at info@wellcomeopenresearch.org.
Image credits
Eimeria tenella: Damer Blake, Royal Veterinary College
Nymphalis polychloros: Will Langdon, University of Oxford
Bufo bufo: Ashleigh Whiffin, National Museums Scotland